Grad student automates gene editing to detect indications of cancer
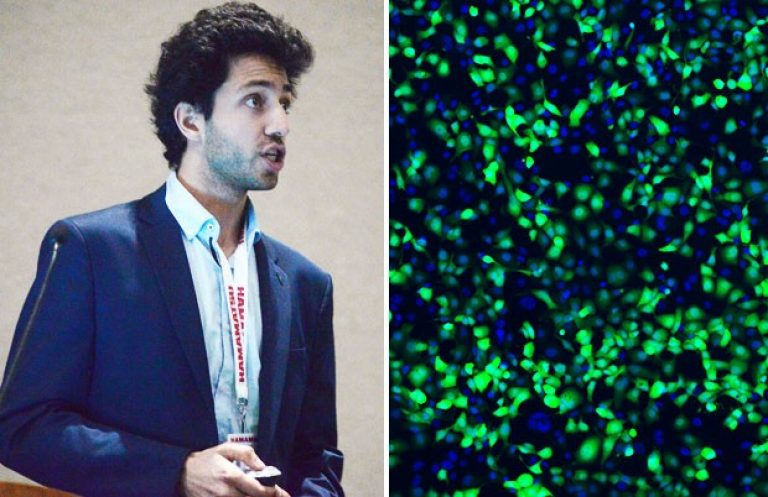
With a BSc in biology tucked under his arm, Concordia student Hugo Sinha took an unusual next step.
“I made the leap to graduate school in the Department of Electrical and Computer Engineering,” says Sinha, who recently returned from presenting his research at the Society for Laboratory Automation and Screening (SLAS) Conference 2018 in San Diego.
At 23 years old, he was the youngest presenter and winner of a Tony B. Academic Travel Award. His work bridges the gap between engineering and biology by automating gene editing for rapid and routine identification of cancer genes.
“I strive to find the coordinates of certain gene 'suspects' thought to be involved in cancer pathways and systematically eliminate those by genome editing,” says Sinha.
Biology, computer science, electrical engineering, genetics — it all factors into Sinha’s multidisciplinary approach. And his results are promising.
First, there’s CRISPR
Working with his mentor, Steve Shih, founder of the Shih Microfluidics Laboratory and assistant professor of electrical and computer engineering in Concordia’s Faculty of Engineering and Computer Science, Sinha designed and validated a new liquid-handling tool to automate gene editing processes specific to CRISPR-Cas9 workflow.
CRISPR, or "Clustered Regularly Interspaced Short Palindromic Repeats,” is a genetic engineering technique that works like a pair of scissors to cut, insert or reorder specific genetic fragments, creating changes in the biological cell to understand gene function.
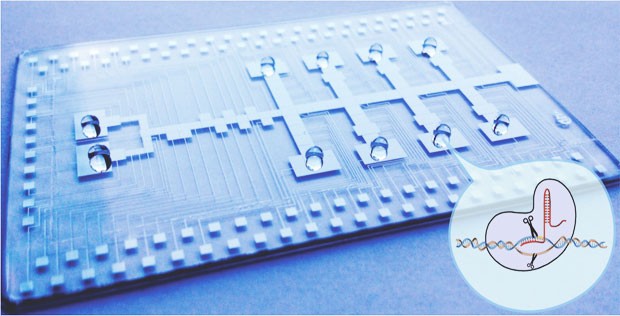 The automated microfluidic tool targets a specific set of genes in lung cancer cells and determines which are modulators of cancer progression. INSERT: A schematic representation of the CRISPR Cas9 process.
The automated microfluidic tool targets a specific set of genes in lung cancer cells and determines which are modulators of cancer progression. INSERT: A schematic representation of the CRISPR Cas9 process.
Automation: faster, better and cheaper
“I’m automating the CRISPR pipeline and making the biology component easier and faster to work with,” says Sinha.
“On this 2-inch by 3-inch platform, I’m able to culture human cells, insert the gene editing material inside the cells — lipid-mediated transfection — and perform cell analysis.”
The automated tool is powered by the manipulation of droplets on an open array of electrodes. This technology, called digital microfluidics, falls into the "lab-on-a-chip" category — pipettes, beakers, flasks and test tubes are all figuratively crammed onto a single hand-held device.
Sinha often thinks about his research as a criminal investigation — CSI Cancer, if you will.
There are plenty of gene "suspects" to sort through or screen. He proceeds by elimination — "knocking-out" specific genes and studying the effect that has on cell proliferation. The CRISPR-Cas9 technology enables researchers to rapidly and precisely eliminate certain genes.
However, this becomes either labor-intensive (if done manually) or expensive (if you need robotics). Automating the CRISPR-Cas9 pipeline for cancer research at a low price is a game-changer.
Proof of concept: lung cancer cells
“The first step of my project and my proof-of-concept is to use my platform to ‘knock out’ the green fluorescent protein gene in lung cancer cells by disrupting it using CRISPR,” says Sinha.
“Looking at microscopy images, we're able to see which ones have been knocked-out (they’re no longer green) and which ones have not (they’re still green).
“From that, we're able to determine the knock-out efficiency, or the number of cells that have been modified. Once this is done at high efficiencies, we know that our platform can effectively tweak genomes and we can move on to target more useful genes, that may be involved in cancer pathways.”
The study demonstrates that Sinha’s genome editing-on-a-chip platform will greatly speed up validation of loss-of-function screens, at relatively low cost, with a minute amount of cell material and without the need for enrichment analysis based on next-generation sequencing profiles as required by pooled screens.
"One of the major challenges related to cancer is to find the genetic culprits,” says Shih. “Hugo has taken a step forward in that he can now quickly find the genetic causes of cancer by removing genes that are possible drivers of cancer.
“This was previously impossible since automation was too expensive. Now with a smaller platform, with cheaper automation, we can tackle this challenge.
Shih says SLAS was the perfect place for Hugo to present his work since it bridges the gap between automation and life sciences. “This means bringing the engineers and biologists together to solve major societal challenges like cancer."
Sinha describes his platform as a “work in progress” at this stage, but he and Shih would like to commercialize it as an affordable bench-top instrument for researchers in many fields.
“I've been talking about cancer a lot, but this strategy can be used for any type of gene editing research,” says Sinha.
“There's a wide range of applications for CRISPR — from generating mutant mosquitos that can't transmit malaria to trying to eradicate HIV. That's why we want to automate this process using our technology. It's easy to reprogram, it’s modular, and it’s easy to integrate into other scientists’ workflow.”
Listen to Shih and Sinha describe their work on Concordia's 2067 podcast series. Learn more about the Shih Microfluidics Laboratory at Concordia.
Find out more about how CRISPR works.